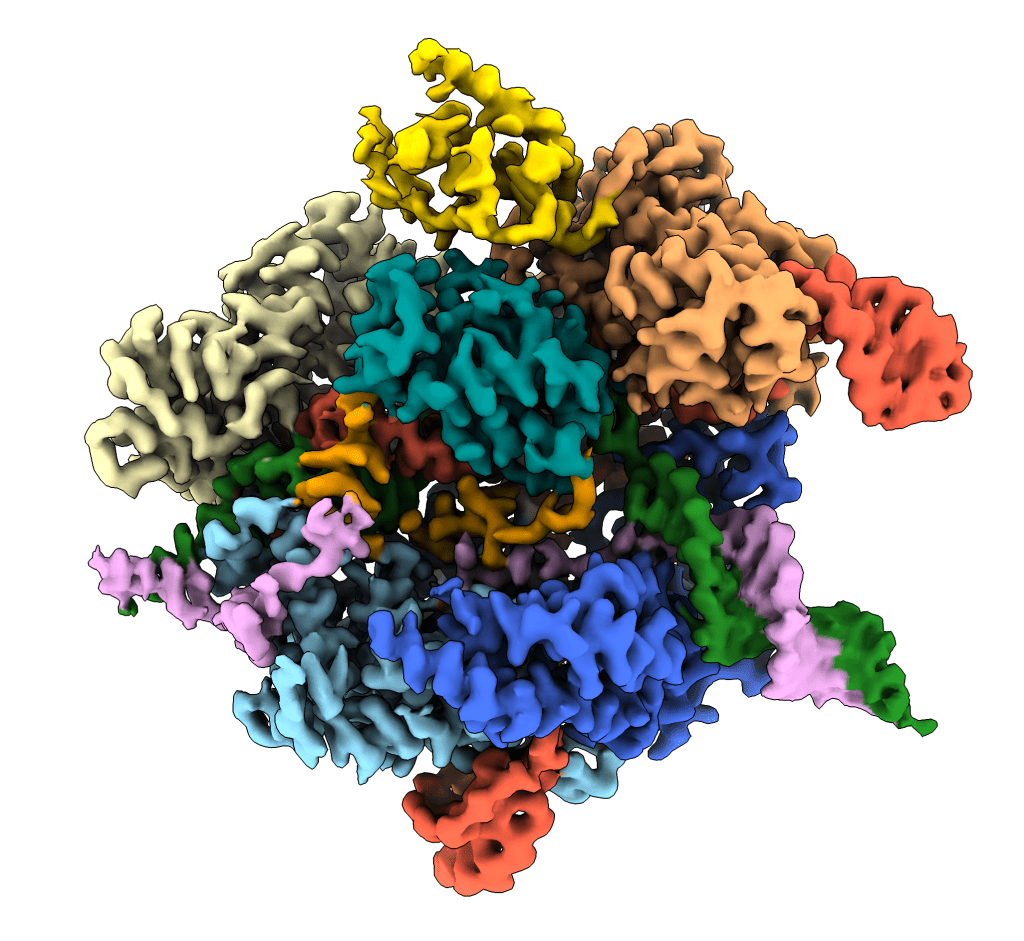
CAS9 RNA-GUIDED ENDONUCLEASES
The widespread use of CRISPR/Cas9 as a programmable genome editing tool has been hindered by off-target DNA cleavage. While analysis of such off-target editing events have enabled the development of Cas9 variants with greater discrimination against mismatches, the underlying molecular mechanisms by which Cas9 rejects or accepts mismatches are poorly understood. We use a combination of kinetic analysis and cryo-EM to interrogate the mechanisms of mismatch surveillance by Cas9. This unique approach allows us to interrogate the different stages of Cas9 activation, from DNA binding to R-loop formation to DNA cutting. This is essential to gain precise molecular insights into the mechanisms that underpin Cas9 activation which will allow us to develop the next generation of high-fidelity Cas9 variants.
See: Gong et al., 2018, Cell Reports; Liu et al., 2020, Nature Communications; Bravo et al., 2022, Nature; Bravo et al., 2022 Current Opinion in Biotechnology

CASCADE SURVEILLANCE COMPLEXES
In bacteria, the clustered regularly interspaced short palindromic repeats (CRISPR)–associated (Cas) DNA-targeting complex Cascade (CRISPR-associated complex for antiviral defense) uses CRISPR RNA (crRNA) guides to bind complementary DNA targets at sites adjacent to a trinucleotide signature sequence called the protospacer adjacent motif (PAM). The Cascade complex then recruits Cas3, a nuclease-helicase that catalyzes unwinding and cleavage of foreign double-stranded DNA (dsDNA) bearing a sequence matching that of the crRNA. We use our favorite tool – cryo-EM – to study the assembly, function and molecular mechanisms of these fascinating complexes.
See: O’Brien et al., 2020, Nature Communications; McBride & Schwartz et al., 2020, Molecular Cell; Schwartz, McBride & Bravo et al., 2022, Nature Communications; O’Brien, Bravo et al., 2023, Molecular Cell; Schwartz, Bravo et al., bioRxiv, 2023
